MRI Image Segmentation in MATLAB
Last Updated :
12 Dec, 2022
In the domain of digital image processing, sometimes we need to separate the main object from the image for clear observation. Image segmentation is the process that enables this partitioning. In this method, each pixel is assigned a label, and pixels that share some characteristics are assigned the same label number. This technique is widely used in the medical domain to locate the object of interest.
It is a technique to partition a digital image into multiple segments. This process is widely used in medical diagnosis. Here in this article, we have used morphological operations to segment the brain part from the MRI image. The segmentation is carried out in order to facilitate the analysis of the segmented images.
What is Medical imaging?
This term refers to the technique which medical professionals use to view inside the human body in order to diagnose, monitor, and treat. There are various techniques available in the modern scientific age.
- MRI: Magnetic resonance imaging
- CT Scan: Computed Tomography Scan
- X-Ray: Using electromagnetic waves called X-rays.
- Ultrasound: Uses sound waves to create pictures of inner body tissues. It does not use any radiation.
Types of Image Segmentations:
There exist many types of image segmentation.
- Thresholding Segmentation
- Edge-Based Segmentation
- Region-Based Segmentation
- Watershed Segmentation
In this article, we are working on separating the brain part of the MRI image.
Morphological operations used:
- Open: It converts small white portions or a bunch of bright pixels into dark portions without changing the size of a larger dark portion.
- Close: It converts small black portions or a bunch of dark pixels into bright portions without changing the size of the larger white portion.
Function Used:
- imread( ) inbuilt-function is used to read the image.
- imtool( ) inbuilt-function is used to display the image.
- graythresh( ) inbuilt-function is used to calculate the Otsu threshold for converting grayscale image into binary.
- im2bw( ) inbuilt-function is used to convert grayscale image into binary.
- strel( ) inbuilt-function is used to define the structuring element.
- imopen( ) inbuilt-function is used to apply opening morphological operation.
- bwlabel( ) inbuilt-function is used to assign the label to each pixel of image.
- imclose( ) inbuilt-function is used to apply closing morphological operation.
- double( ) inbuilt-function is used to convert unit8 object into double format.
Steps:
- Read the MRI image.
- Convert it into binary.
- Apply opening operation.
- Apply connected component analysis.
- Identify the brain part and separate it out.
- Display different images in intermediate steps.
Example:
Matlab
k=imread("mrii.jpg");
imtool(k,[]);
k1=im2bw(k,graythresh(k));
imtool(k1);
SE=strel('disk',7,4);
k2=imopen(k1,SE);
imtool(k2);
b=bwlabel(k2);
imtool(b,[]);
b(b~=9)=0;
imtool(b);
k3=imclose(b,strel('disk',18));
imtool(k3);
k4=k3.*double(k);
imtool(k4,[]);
|
Output:
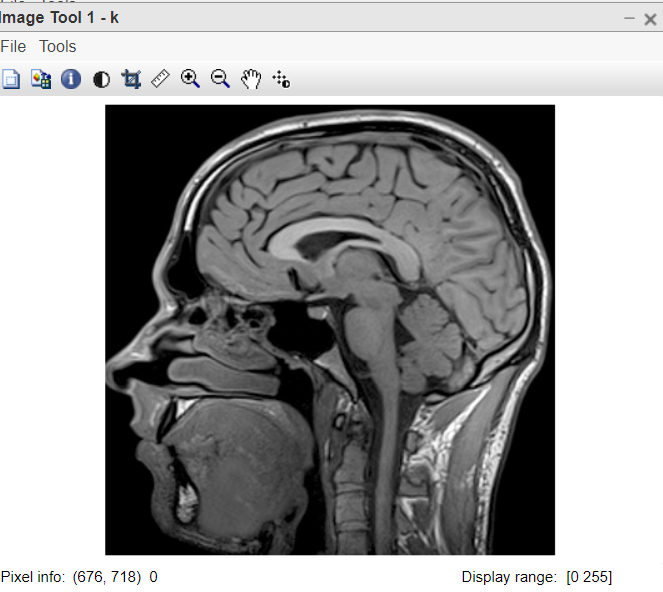
Figure 1: Original image
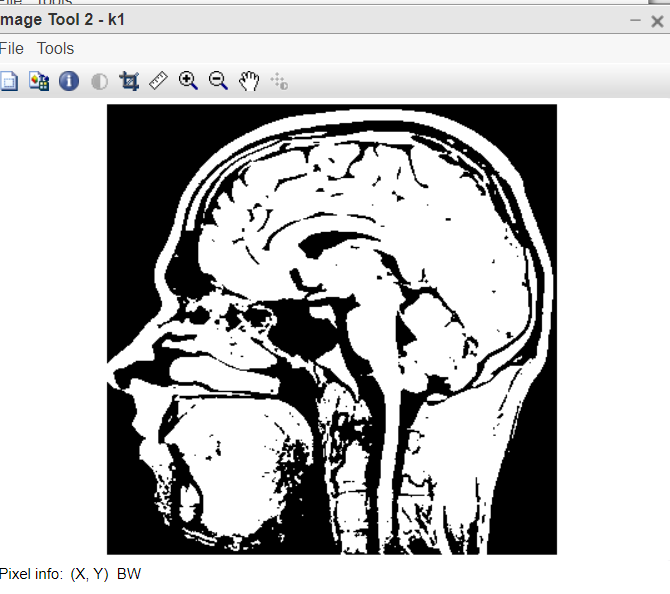
Figure 2: Binary image
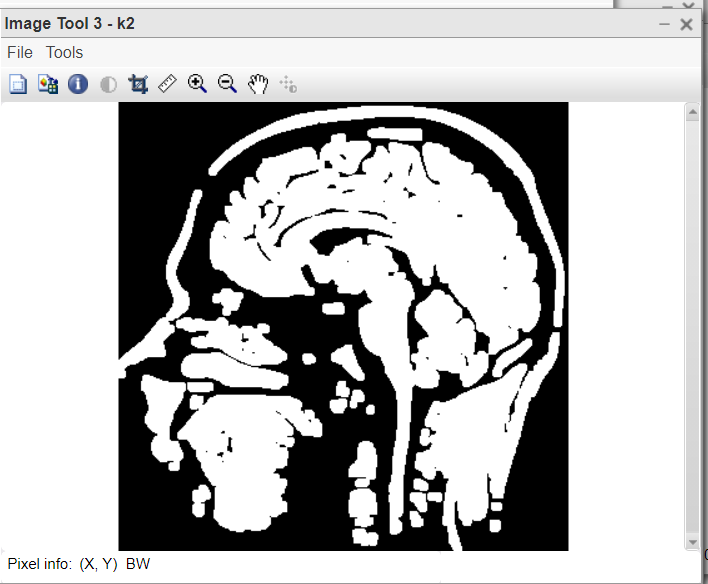
Figure 3:After applying to open
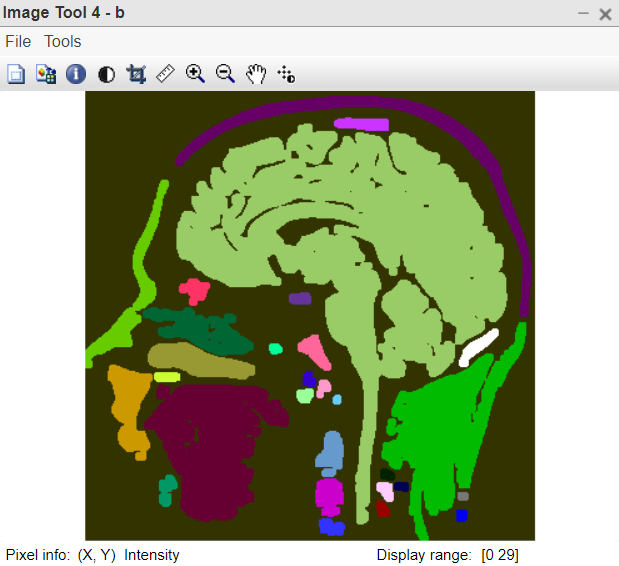
Figure 4:ColourMap image
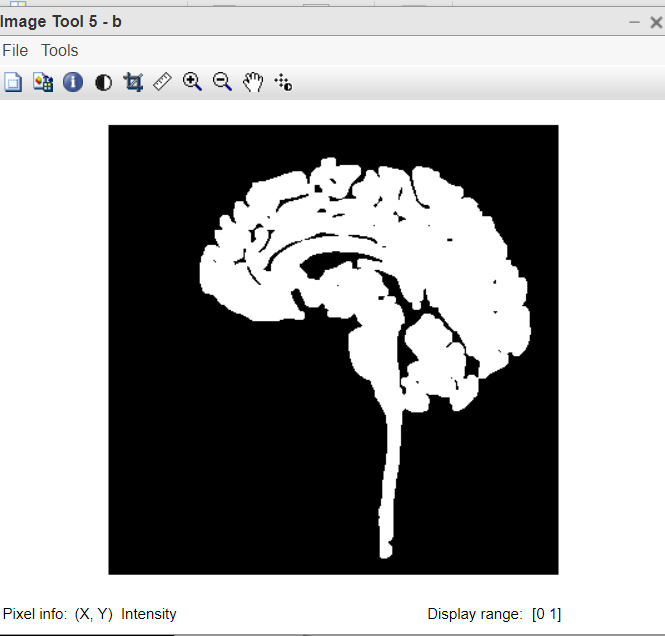
Figure 5:Biggest connected component
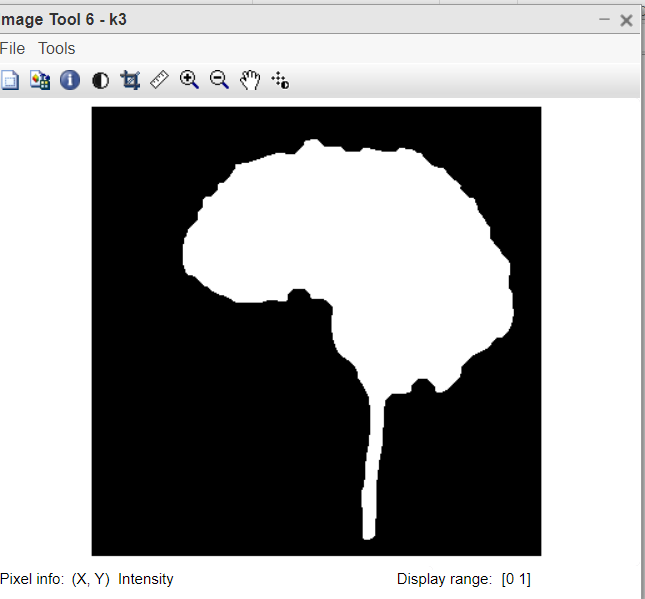
Figure 6: After applying to close on the brain part
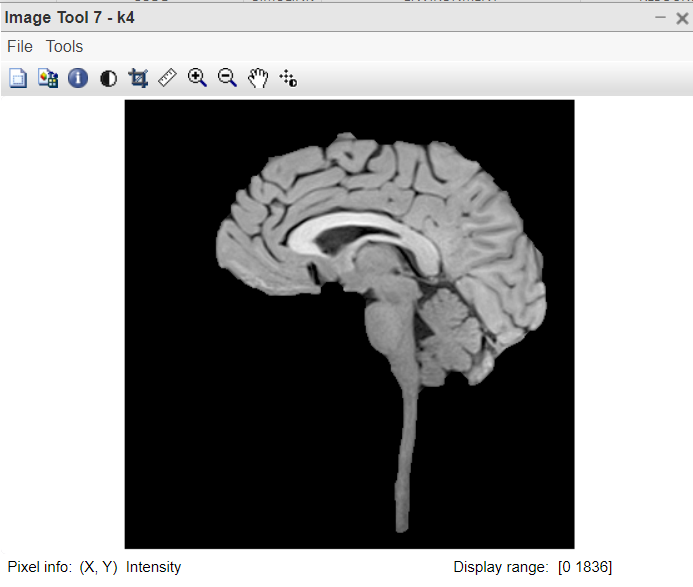
Figure 7:Brain extracted from the original image
Code explanation:
- k=imread(“mrii.jpg”); this line reads the mri image
- k1=im2bw(k,graythresh(k)); this line converts image into binary image
- SE=strel(‘disk’,7,4); this line defines the structuring element.
- k2=imopen(k1,SE); this line applies the opening operation
- b=bwlabel(k2); this line applies connected component analysis
- b(b~=9)=0; this line sets all other component as 0 except brain, since brain is labeled as 9.
- k3=imclose(b,strel(‘disk’,18)); this line closes the black pixels inside brain part for Inside the brain part, black portion is there.
- k4=k3.*double(k); this line extracts the brain from original image
Share your thoughts in the comments
Please Login to comment...