Bagging and Random Forest for Imbalanced Classification
Last Updated :
11 Mar, 2024
Ensemble learning techniques like bagging and random forests have gained prominence for their effectiveness in handling imbalanced classification problems. In this article, we will delve into these techniques and explore their applications in mitigating the impact of class imbalance.
Classification problems are fundamental to machine learning and find applications across various domains. However, a prevalent issue in such tasks is handling imbalanced datasets, where one class significantly outweighs the other. This class imbalance presents a hurdle for conventional classifiers as they often exhibit a bias toward the majority class, resulting in skewed models.
Bagging for Imbalanced Classification
Let us start by exploring Bagging Techniques for Imbalanced Classification problem.
Standard Bagging
Ensemble learning is a machine learning approach that involves using multiple learning algorithms to create a stronger model than an individual model. Bagging, or bootstrap aggregating, is one of these techniques involving creation of multiple models on different subsets of the training data and then combining their predictions. It combines two key concepts: bootstrapping and aggregation.
- Bootstrap: Creating multiple datasets through sampling with replacement from the original dataset. This technique allows for the creation of multiple “bootstrapped” datasets that are similar but not identical to the original dataset.
- Aggregation: Once the bootstrapped datasets are created, an algorithm (such as a decision tree) is trained on each dataset. The predictions from each model are then aggregated or combined to make the final prediction. This aggregation helps improve the overall accuracy of the model.
Implementation
We will explore the impact of bagging on imbalanced classification using a simplified example on an imbalanced dataset using the scikit-learn library. For this, we generate an imbalanced dataset with 2 target classes and a class distribution of 90% for the majority class and 10% for the minority class.
Python
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
X, y = make_classification(n_samples=1500, n_features=15, n_informative=5, n_redundant=1, n_classes=2, weights=[0.90, 0.10])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
|
Let’s visualize this class distribution:
Python
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
label_counts = np.bincount(y)
fig, ax = plt.subplots(figsize=(8, 6))
ax = sns.barplot(x=np.arange(2), y=label_counts, palette="Set1")
ax.set_xticks(np.arange(2))
ax.set_xticklabels(['Class 1', 'Class 2'])
ax.set_title("Count Plot of Synthetic Datapoints", fontsize=16)
ax.set_xlabel("Classes", fontsize=14)
ax.set_ylabel("# Samples", fontsize=14)
plt.show()
|
Output:
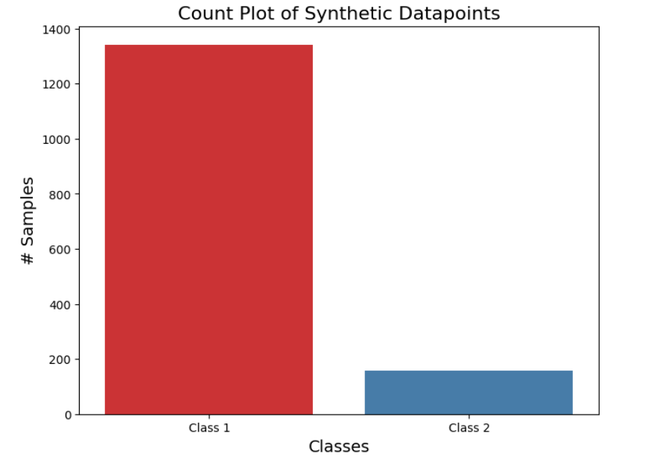
After visualization we will train our Bagging Model:
Python
from sklearn.ensemble import BaggingClassifier
from sklearn.metrics import accuracy_score
bagging_clf = BaggingClassifier()
bagging_clf.fit(X_train, y_train)
y_pred = bagging_clf.predict(X_test)
acc_bag = accuracy_score(y_test, y_pred)
print(" Bagging Classifier - Test Accuracy:", round(acc_bag, 2))
|
Output:
Bagging Classifier - Test Accuracy: 0.92
Bagging With Random Undersampling
In order to further improve the performance of Bagging Classifier on Imbalanced Dataset, we can perform random undersampling of the majority class. This technique involves randomly eliminating instances from the majority class to achieve a balanced distribution in each bootstrapped sample. By doing so, the model is less likely to exhibit bias towards the majority class, thus enhancing its accuracy in classifying minority instances.
Implementation
Python
from sklearn.ensemble import BaggingClassifier
from imblearn.under_sampling import RandomUnderSampler
from sklearn.metrics import accuracy_score
rus = RandomUnderSampler()
X_train_resampled, y_train_resampled = rus.fit_resample(X_train, y_train)
bagging_classifier = BaggingClassifier()
bagging_classifier.fit(X_train_resampled, y_train_resampled)
y_pred = bagging_classifier.predict(X_test)
acc_rndm = accuracy_score(y_test, y_pred)
print(" Bagging Classifier with Random Undersampling - Test Accuracy:", round(acc_rndm, 2))
|
Output:
Bagging Classifier with Random Undersampling - Test Accuracy: 0.96
Random Forest for Imbalanced Classification
Now that we have an idea about Bagging techniques, let’s understand Random Forest and how it can be used for Imbalanced Classification problem.
Standard Random Forest
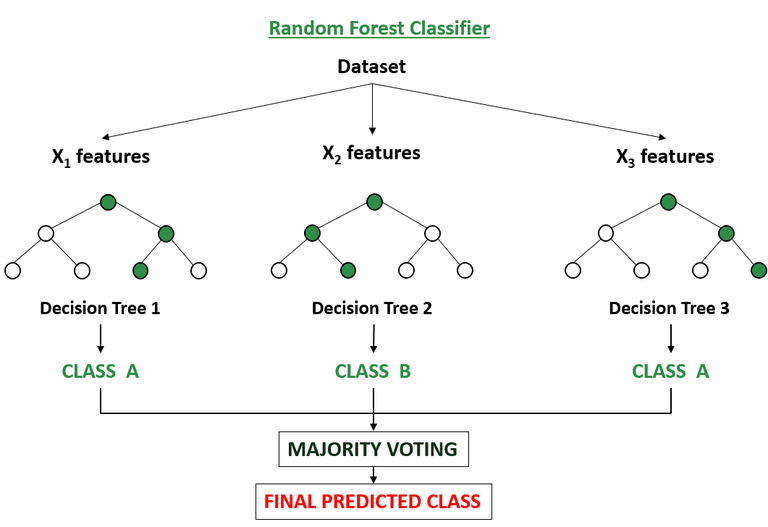
Random forest is a variation of bagging that deploys decision trees as the base models. It trains each decision tree on a random subset of the training data and a random subset of the features and at the time of prediction, the random forest algorithm aggregates the predictions of all the individual decision trees to make a final prediction with higher predictive accuracy and also control over-fitting.
For classification tasks, after each tree has been trained on a subset of the training data, in order to make a prediction for a new data point, the random forest algorithm passes the data point through each decision tree in the forest. After this, each tree independently predicts the class of the data point. If a certain class receives the highest number of votes across all trees, that class becomes the predicted class.
Implementation
Now we will go through a practical example to explore the impact of random forest on imbalanced classification.
Python
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
X, y = make_classification(n_samples=1500, n_features=15, n_informative=5, n_redundant=1, n_classes=2, weights=[0.90, 0.10])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
|
After data split we will train Random Forest model:
Python
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
rf = RandomForestClassifier(n_estimators=100)
rf.fit(X_train, y_train)
y_pred_rf = rf.predict(X_test)
acc_rf = accuracy_score(y_test, y_pred_rf)
print(f"Standard Random Forest Accuracy: {acc_rf:.2f}")
|
Output:
Standard Random Forest Accuracy: 0.96
Random Forest With Class Weighting
Another way that we can handle class imbalance is by adjusting the weights assigned to each class in the Random Forest algorithm aka Class Weighting. When we give more weight to the minority class and less weight to the majority class, the algorithm pays more attention to the minority class during training. This allows the model to learn from the imbalanced data more effectively, leading to better performance on predicting the minority class.
Implementation
Python
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
rf_cw = RandomForestClassifier(random_state=42, class_weight="balanced")
rf_cw.fit(X_train, y_train)
y_pred_cw = rf_cw.predict(X_test)
acc_cw = accuracy_score(y_test, y_pred_cw)
print(f"Random Forest with Class Weighting Accuracy: {acc_cw:.2f}")
|
Output:
Random Forest with Class Weighting Accuracy: 0.93
Random Forest With Bootstrap Class Weighting
Bootstrapping process is a vital aspect of Random Forests, and by combining Class Weighting with bootstrap we can quite effectively handle class imbalance in our data. In a traditional Random Forest model, each tree is trained on a bootstrap sample of the original data set, where samples may be repeated or left out. In Random Forest with bootstrap class weighting, additional weights based on class distribution are given to samples during bootstrapping, focusing more on minority class samples for improved classification accuracy on minority class instances.
Implementation
Python
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
rf_bcw = RandomForestClassifier(random_state=42, class_weight='balanced_subsample')
rf_bcw.fit(X_train, y_train)
y_pred_bcw = rf_bcw.predict(X_test)
acc_bcw = accuracy_score(y_test, y_pred_bcw)
print(f"Random Forest with Bootstrap Class Weighting Accuracy: {acc_bcw:.2f}")
|
Output:
Random Forest with Bootstrap Class Weighting Accuracy: 0.94
Random Forest With Random Undersampling
Similar to bagging, random undersampling can be combined with random forests for improved performance on imbalanced datasets. By randomly removing some instances from the majority class in each bootstrap sample, we can balance the class distribution. This allows random forest classifiers to focus more on the minority class during training, potentially improving their ability to learn effectively from minority class instances and classify them correctly.
Implementation
Python
from sklearn.ensemble import RandomForestClassifier
from imblearn.under_sampling import RandomUnderSampler
from sklearn.metrics import accuracy_score
rus = RandomUnderSampler(random_state=42)
X_train_resampled, y_train_resampled = rus.fit_resample(X_train, y_train)
rf_rus = RandomForestClassifier(random_state=42)
rf_rus.fit(X_train_resampled, y_train_resampled)
y_pred_rus = rf_rus.predict(X_test)
acc_rus = accuracy_score(y_test, y_pred_rus)
print(f"Random Forest with Random Undersampling Accuracy: {acc_rus:.2f}")
|
Output:
Random Forest with Random Undersampling Accuracy: 0.89
Conclusion
In conclusion, ensemble learning techniques such as bagging and random forests offer effective solutions to the challenges posed by imbalanced classification problems. By combining multiple base classifiers these techniques can improve model performance and generalization on imbalanced datasets. Through standard bagging, bagging with random undersampling, and various adaptations of random forests, practitioners can address class imbalance and build more robust classification models.
Share your thoughts in the comments
Please Login to comment...