The process of splitting images into multiple layers, represented by a smart, pixel-wise mask is known as Image Segmentation. It involves merging, blocking, and separating an image from its integration level. Splitting a picture into a collection of Image Objects with comparable properties is the first stage in image processing. Scikit-Image is the most popular tool/module for image processing in Python.
Installation
To install this module type the below command in the terminal.
pip install scikit-image
Converting Image Format
RGB to Grayscale
rgb2gray module of skimage package is used to convert a 3-channel RGB Image to one channel monochrome image. In order to apply filters and other processing techniques, the expected input is a two-dimensional vector i.e. a monochrome image.
skimage.color.rgb2gray() function is used to convert an RGB image to Grayscale format
Syntax : skimage.color.rgb2gray(image)
Parameters : image : An image – RGB format
Return : The image – Grayscale format
Code:
Python3
from skimage import data
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
plt.figure(figsize=(15, 15))
coffee = data.coffee()
plt.subplot(1, 2, 1)
plt.imshow(coffee)
gray_coffee = rgb2gray(coffee)
plt.subplot(1, 2, 2)
plt.imshow(gray_coffee, cmap="gray")
|
Output:
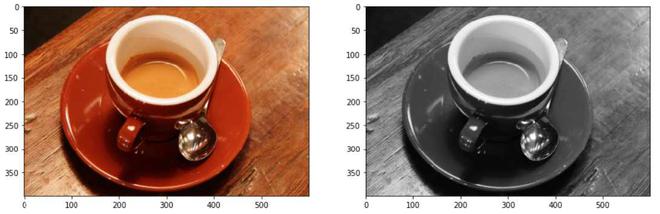

Converting 3-channel image data to 1-channel image data
Explanation: By using rgb2gray() function, the 3-channel RGB image of shape (400, 600, 3) is converted to a single-channel monochromatic image of shape (400, 300). We will be using grayscale images for the proper implementation of thresholding functions. The average of the red, green, and blue pixel values for each pixel to get the grayscale value is a simple approach to convert a color picture 3D array to a grayscale 2D array. This creates an acceptable gray approximation by combining the lightness or brightness contributions of each color band.
The HSV (Hue, Saturation, Value) color model remaps the RGB basic colors into dimensions that are simpler to comprehend for humans. The RGB color space describes the proportions of red, green, and blue in a colour. In the HSV color system, colors are defined in terms of Hue, Saturation, and Value.
skimage.color.rgb2hsv() function is used to convert an RGB image to HSV format
Syntax : skimage.color.rgb2hsv(image)
Parameters : image : An image – RGB format
Return : The image – HSV format
Code:
Python3
from skimage import data
from skimage.color import rgb2hsv
import matplotlib.pyplot as plt
plt.figure(figsize=(15, 15))
coffee = data.coffee()
plt.subplot(1, 2, 1)
plt.imshow(coffee)
hsv_coffee = rgb2hsv(coffee)
plt.subplot(1, 2, 2)
hsv_coffee_colorbar = plt.imshow(hsv_coffee)
plt.colorbar(hsv_coffee_colorbar, fraction=0.046, pad=0.04)
|
Output:
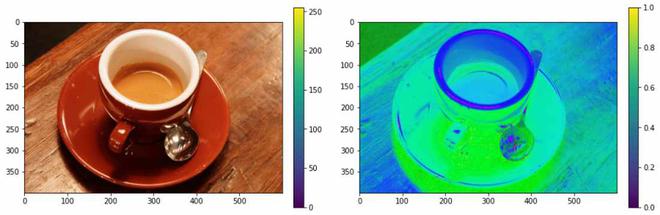
Converting the RGB color format to HSV color format
Supervised Segmentation
For this type of segmentation to proceed, it requires external input. This includes things like setting a threshold, converting formats, and correcting external biases.
Segmentation by Thresholding – Manual Input
An external pixel value ranging from 0 to 255 is used to separate the picture from the background. This results in a modified picture that is larger or less than the specified threshold.
Python3
from skimage import data
from skimage import filters
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
coffee = data.coffee()
gray_coffee = rgb2gray(coffee)
plt.figure(figsize=(15, 15))
for i in range(10):
binarized_gray = (gray_coffee > i*0.1)*1
plt.subplot(5,2,i+1)
plt.title("Threshold: >"+str(round(i*0.1,1)))
plt.imshow(binarized_gray, cmap = 'gray')
plt.tight_layout()
|
Output:

Explanation: The first step in this thresholding is implemented by normalizing an image from 0 – 255 to 0 – 1. A threshold value is fixed and on the comparison, if evaluated to be true, then we store the result as 1, otherwise 0. This globally binarized image can be used to detect edges as well as analyze contrast and color difference.
Segmentation by Thresholding Using skimage.filters module
The Niblack and Sauvola thresholding technique is specifically developed to improve the quality of microscopic images. It’s a local thresholding approach that changes the threshold depending on the local mean and standard deviation for each pixel in a sliding window. Otsu’s thresholding technique works by iterating over all possible threshold values and computing a measure of dispersion for the sample points on either side of the threshold, i.e. either in foreground or background. The goal is to determine the smallest foreground and background spreads possible.
skimage.filters.threshold_otsu() function is used to return threshold value based on Otsu’s method.
Syntax : skimage.filters.threshold_otsu(image)
Parameters :
- image : An image – Monochrome format
- nbins : Number of bins required for histogram calculation
- hist : Histogram from which threshold has to be calculated
Return : threshold : Larger pixel intensity
skimage.filters.threshold_niblack() function is a local thresholding function that returns a threshold value for every pixel based on Niblack’s method.
Syntax : skimage.filters.threshold_niblack(image)
Parameters :
- image : An image – Monochrome format
- window_size : Window size – odd integer
- k : A positive parameter
Return : threshold : A threshold mask equal to the shape of the image
skimage.filters.threshold_sauvola() function is a local thresholding function that returns a threshold value for every pixel based on Sauvola’s method.
Syntax : skimage.filters.threshold_sauvola(image)
Parameters :
- image : An image – Monochrome format
- window_size : Window size – odd integer
- k : A positive parameter
- r : A positive parameter – dynamic range of standard deviation
Return : threshold : A threshold mask equal to the shape of the image
Code:
Python3
from skimage import data
from skimage import filters
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
plt.figure(figsize=(15, 15))
coffee = data.coffee()
gray_coffee = rgb2gray(coffee)
threshold = filters.threshold_otsu(gray_coffee)
binarized_coffee = (gray_coffee > threshold)*1
plt.subplot(2,2,1)
plt.title("Threshold: >"+str(threshold))
plt.imshow(binarized_coffee, cmap = "gray")
threshold = filters.threshold_niblack(gray_coffee)
binarized_coffee = (gray_coffee > threshold)*1
plt.subplot(2,2,2)
plt.title("Niblack Thresholding")
plt.imshow(binarized_coffee, cmap = "gray")
threshold = filters.threshold_sauvola(gray_coffee)
plt.subplot(2,2,3)
plt.title("Sauvola Thresholding")
plt.imshow(threshold, cmap = "gray")
binarized_coffee = (gray_coffee > threshold)*1
plt.subplot(2,2,4)
plt.title("Sauvola Thresholding - Converting to 0's and 1's")
plt.imshow(binarized_coffee, cmap = "gray")
|
Output

Explanation: These local thresholding techniques use mean and standard deviation as their primary computational parameters. Their final local pixel value is felicitated by other positive parameters too. This is done to ensure the separation between the object and the background.
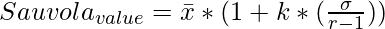
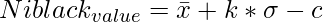
where \bar x and \sigma represents mean and standard deviation of the pixel intensities respectively.
Active Contour Segmentation
The concept of energy functional reduction underpins the active contour method. An active contour is a segmentation approach that uses energy forces and restrictions to separate the pixels of interest from the remainder of the picture for further processing and analysis. The term “active contour” refers to a model in the segmentation process.
skimage.segmentation.active_contour() function active contours by fitting snakes to image features
Syntax : skimage.segmentation.active_contour(image, snake)
Parameters :
- image : An image
- snake : Initial snake coordinates – for bounding the feature
- alpha : Snake length shape
- beta : Snake smoothness shape
- w_line : Controls attraction – Brightness
- w_edge : Controls attraction – Edges
- gamma : Explicit time step
Return : snake : Optimised snake with input parameter’s size
Code:
Python3
import numpy as np
import matplotlib.pyplot as plt
from skimage.color import rgb2gray
from skimage import data
from skimage.filters import gaussian
from skimage.segmentation import active_contour
astronaut = data.astronaut()
gray_astronaut = rgb2gray(astronaut)
gray_astronaut_noiseless = gaussian(gray_astronaut, 1)
x1 = 220 + 100*np.cos(np.linspace(0, 2*np.pi, 500))
x2 = 100 + 100*np.sin(np.linspace(0, 2*np.pi, 500))
snake = np.array([x1, x2]).T
astronaut_snake = active_contour(gray_astronaut_noiseless,
snake)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
ax.imshow(gray_astronaut_noiseless)
ax.plot(astronaut_snake[:, 0],
astronaut_snake[:, 1],
'-b', lw=5)
ax.plot(snake[:, 0], snake[:, 1], '--r', lw=5)
|
Output:

Explanation: The active contour model is among the dynamic approaches in image segmentation that uses the image’s energy restrictions and pressures to separate regions of interest. For segmentation, an active contour establishes a different border or curvature for each section of the target object. The active contour model is a technique for minimizing the energy function resulting from external and internal forces. An exterior force is specified as curves or surfaces, while an interior force is defined as picture data. The external force is a force that allows initial outlines to automatically transform into the forms of objects in pictures.
Chan-Vese Segmentation
The well-known Chan-Vese iterative segmentation method splits a picture into two groups with the lowest intra-class variance. This algorithm uses sets that are iteratively evolved to minimize energy, which is characterized by weights corresponding to the total of variations in intensity from the overall average outside the segmented region, the sum of differences from the overall average within the feature vector, and a term that is directly proportional to the length of the fragmented region’s edge.
skimage.segmentation.chan_vese() function is used to segment objects using the Chan-Vese Algorithm whose boundaries are not clearly defined.
Syntax : skimage.segmentation.chan_vese(image)
Parameters :
- image : An image
- mu : Weight – Edge Length
- lambda1 : Weight – Difference from average
- tol : Tolerance of Level set variation
- max_num_iter : Maximum number of iterations
- extended_output : Tuple of 3 values is returned
Return :
- segmentation : Segmented Image
- phi : Final level set
- energies: Shows the evolution of the energy
Code:
Python3
import matplotlib.pyplot as plt
from skimage.color import rgb2gray
from skimage import data, img_as_float
from skimage.segmentation import chan_vese
fig, axes = plt.subplots(1, 3, figsize=(10, 10))
astronaut = data.astronaut()
gray_astronaut = rgb2gray(astronaut)
chanvese_gray_astronaut = chan_vese(gray_astronaut,
max_iter=100,
extended_output=True)
ax = axes.flatten()
ax[0].imshow(gray_astronaut, cmap="gray")
ax[0].set_title("Original Image")
ax[1].imshow(chanvese_gray_astronaut[0], cmap="gray")
title = "Chan-Vese segmentation - {} iterations".
format(len(chanvese_gray_astronaut[2]))
ax[1].set_title(title)
ax[2].imshow(chanvese_gray_astronaut[1], cmap="gray")
ax[2].set_title("Final Level Set")
plt.show()
|
Output:

Explanation: The Chan-Vese model for active contours is a strong and versatile approach for segmenting a wide range of pictures, including some that would be difficult to segment using “traditional” methods such as thresholding or gradient-based methods. This model is commonly used in medical imaging, particularly for brain, heart, and trachea segmentation. The model is based on an energy minimization issue that may be recast in a level set formulation to make the problem easier to solve.
Unsupervised Segmentation
Mark Boundaries
This technique produces an image with highlighted borders between labeled areas, where the pictures were segmented using the SLIC method.
skimage.segmentation.mark_boundaries() function is to return image with boundaries between labeled regions.
Syntax : skimage.segmentation.mark_boundaries(image)
Parameters :
- image : An image
- label_img : Label array with marked regions
- color : RGB color of boundaries
- outline_color : RGB color of surrounding boundaries
Return : marked : An image with boundaries are marked
Code:
Python3
from skimage.segmentation import slic, mark_boundaries
from skimage.data import astronaut
plt.figure(figsize=(15, 15))
astronaut = astronaut()
astronaut_segments = slic(astronaut,
n_segments=100,
compactness=1)
plt.subplot(1, 2, 1)
plt.imshow(astronaut)
plt.subplot(1, 2, 2)
plt.imshow(mark_boundaries(astronaut, astronaut_segments))
|
Output:

Explanation: We cluster the image into 100 segments with compactness = 1 and this segmented image will act as a labeled array for the mark_boundaries() function. Each segment of the clustered image is differentiated by an integer value and the result of mark_boundaries is the superimposed boundaries between the labels.
Simple Linear Iterative Clustering
By combining pixels in the image plane based on their color similarity and proximity, this method generates superpixels. Simple Linear Iterative Clustering is the most up-to-date approach for segmenting superpixels, and it takes very little computing power. In a nutshell, the technique clusters pixels in a five-dimensional color and picture plane space to create small, nearly uniform superpixels.
skimage.segmentation.slic() function is used to segment image using k-means clustering.
Syntax : skimage.segmentation.slic(image)
Parameters :
- image : An image
- n_segments : Number of labels
- compactness : Balances color and space proximity.
- max_num_iter : Maximum number of iterations
Return : labels: Integer mask indicating segment labels.
Code:
Python3
from skimage.segmentation import slic
from skimage.data import astronaut
from skimage.color import label2rgb
plt.figure(figsize=(15,15))
astronaut = astronaut()
astronaut_segments = slic(astronaut,
n_segments=50,
compactness=10)
plt.subplot(1,2,1)
plt.imshow(astronaut)
plt.subplot(1,2,2)
plt.imshow(label2rgb(astronaut_segments,
astronaut,
kind = 'avg'))
|
Output:

Explanation: This technique creates superpixels by grouping pixels in the picture plane based on their color similarity and closeness. This is done in 5-D space, where XY is the pixel location. Because the greatest possible distance between two colors in CIELAB space is restricted, but the spatial distance on the XY plane is dependent on the picture size, we must normalize the spatial distances in order to apply the Euclidean distance in this 5D space. As a result, a new distance measure that takes superpixel size into account was created to cluster pixels in this 5D space.
Felzenszwalb’s Segmentation
Felsenszwalb’s efficient graph-based picture segmentation is computed. It produces an over-segmentation of an RGB picture on the image grid using a quick, minimal spanning tree-based clustering. This may be used to isolate features and identify edges. This algorithm uses the Euclidean distance between pixels. skimage.segmentation.felzenszwalb() function is used to compute Felsenszwalb’s efficient graph-based image segmentation.
Syntax : skimage.segmentation.felzenszwalb(image)
Parameters :
- image : An input image
- scale : Higher value – larger clusters
- sigma : Width of Gaussian kernel
- min_size : Minimum component size
Return : segment_mask : Integer mask indicating segment labels.
Code:
Python3
from skimage.segmentation import felzenszwalb
from skimage.color import label2rgb
from skimage.data import astronaut
plt.figure(figsize=(15,15))
astronaut = astronaut()
astronaut_segments = felzenszwalb(astronaut,
scale = 2,
sigma=5,
min_size=100)
plt.subplot(1,2,1)
plt.imshow(astronaut)
plt.subplot(1,2,2)
plt.imshow(mark_boundaries(astronaut,
astronaut_segments))
|
Output:

With mark_boundaries() function
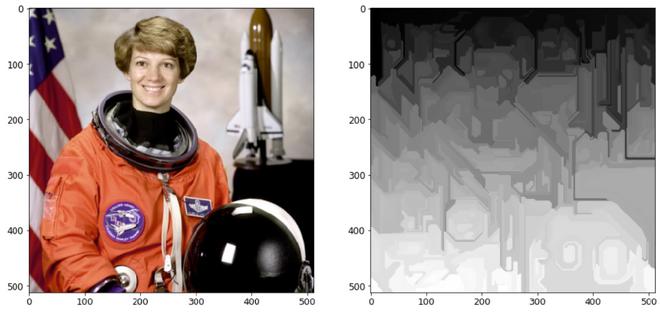
Without mark_boundaries() function
Explanation: Using a rapid, minimal tree structure-based clustering on the picture grid, creates an over-segmentation of a multichannel image. The parameter scale determines the level of observation. Less and larger parts are associated with a greater scale. The diameter of a Gaussian kernel is sigma, which is used to smooth the picture before segmentation. Scale is the sole way to control the quantity of generated segments as well as their size. The size of individual segments within a picture might change dramatically depending on local contrast.
There are many other supervised and unsupervised image segmentation techniques. This can be useful in confining individual features, foreground isolation, noise reduction, and can be useful to analyze an image more intuitively. It is a good practice for images to be segmented before building a neural network model in order to yield effective results.
Like Article
Suggest improvement
Share your thoughts in the comments
Please Login to comment...