Crossover in Genetic Algorithm
Last Updated :
10 Mar, 2023
Crossover is a genetic operator used to vary the programming of a chromosome or chromosomes from one generation to the next. Crossover is sexual reproduction. Two strings are picked from the mating pool at random to crossover in order to produce superior offspring. The method chosen depends on the Encoding Method.
Crossover mask: The choice of which parent contributes to the bit position fI the offspring is given by an additional string called crossover mask similar to bit masks in unity game engine.
Different types of crossover :
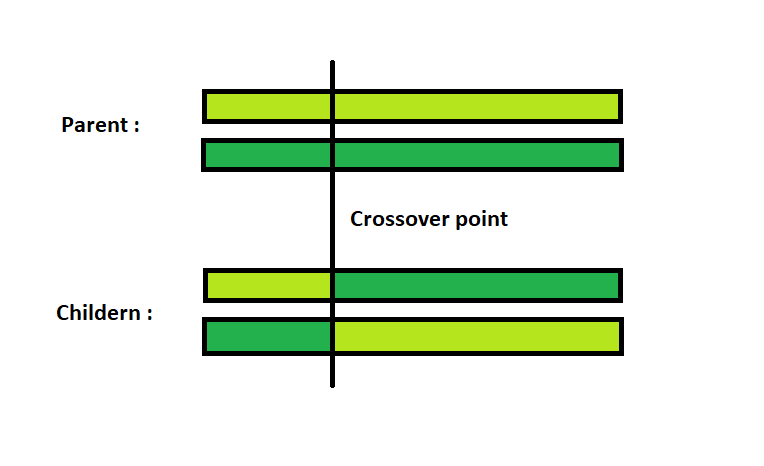
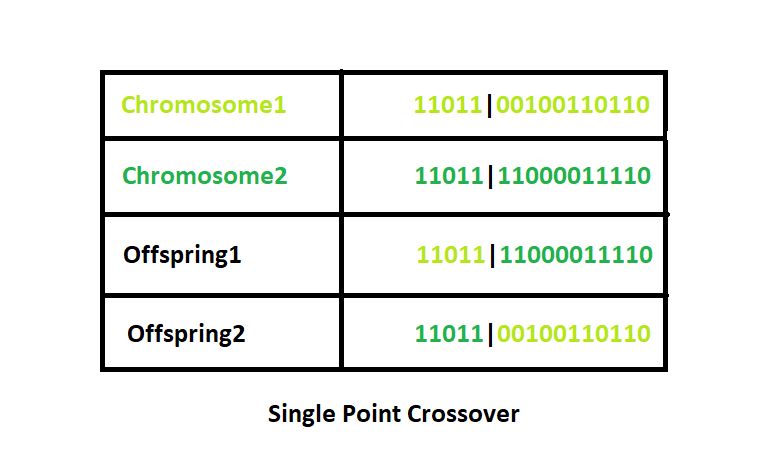
Single Point Crossover: A crossover point on the parent organism string is selected. All data beyond that point in the organism string is swapped between the two parent organisms. Strings are characterized by Positional Bias.
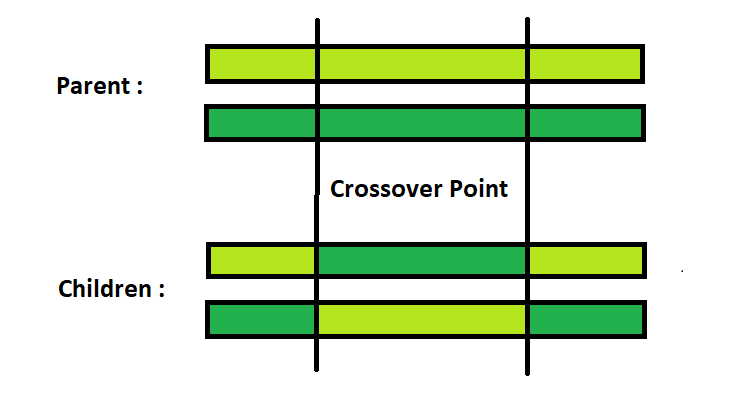
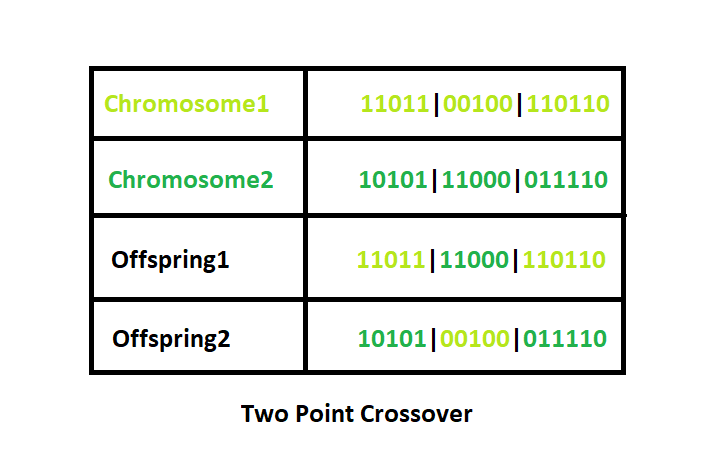
Two-Point Crossover : This is a specific case of a N-point Crossover technique. Two random points are chosen on the individual chromosomes (strings) and the genetic material is exchanged at these points.
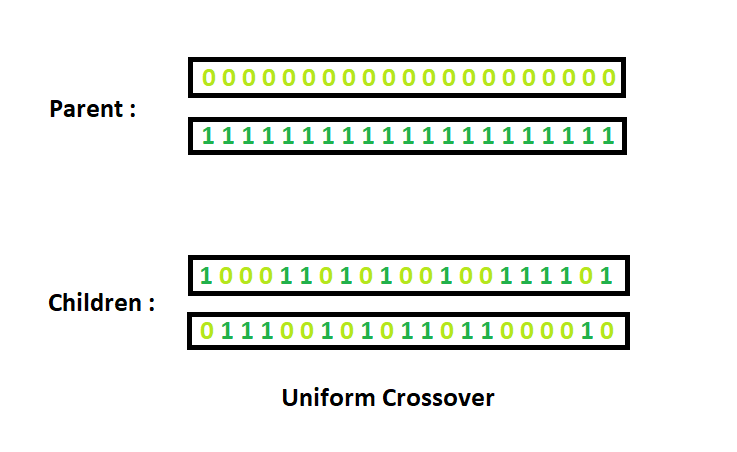
Uniform Crossover: Each gene (bit) is selected randomly from one of the corresponding genes of the parent chromosomes.
Use tossing of a coin as an example technique.
The crossover between two good solutions may not always yield a better or as good a solution. Since parents are good, the probability of the child being good is high. If offspring is not good (poor solution), it will be removed in the next iteration during “Selection”.
Problems with Crossover:
- Depending on coding, simple crossovers can have a high chance to produce illegal offspring.
E.g. in TSP with simple binary or path coding, most offspring will be illegal because not all cities will be in the offspring and some cities will be there more than once.
- Uniform crossover can often be modified to avoid this problem
E.g. in TSP with simple path coding:
Where the mask is 1, copy cities from one parent
Where the mask is 0, choose the remaining cities in the order of the other parent.
Like Article
Suggest improvement
Share your thoughts in the comments
Please Login to comment...